

The findings of a new fuels forecast developed by Wärtsilä over the year highlight growing demand for fuel flexibility in power technologies and across the lifecycle of assets. The project will help to define Wärtsilä's technology development roadmap in the marine and energy segments. For end users the findings offer important insights into the choices they will need to make very soon.
As climate action is driven by international agreements and policies that incentivise investments in clean energy, the first step of the project was to examine the status and progress of these ambitions. According to Mikael Wideskog, Director of Sustainable Fuels & Decarbonisation, Wärtsilä Marine Power and co-chair of the fuel strategy steering group, it is clear that the world is “very far away” from meeting the Paris Agreement target of keeping post-industrial global warming below 2°C.
A report published by the UN Environment Programme in 2019 noted that a 7.6% reduction in emissions is needed every year between 2019 and 2030 to meet that target. By comparison, GHG emissions have been climbing by more than 1.5% a year for the past decade.
Climate urgency
But the transition is underway. Several countries have set timeframes to reach carbon neutrality. Shipping, which is not covered by the Paris Agreement, is working towards ambitions set by the International Maritime Organization to cut total GHG emissions by at least half by 2050, and to zero as soon as possible after that.
Cutting carbon will demand new fuels. There is an emerging realisation that most if not all of these fuels will be derived from biomass and renewably produced hydrogen. According to Wideskog, Wärtsilä’s review of policies and investments reveal that the hydrogen economy is no longer a dream. The EU is a good example. The bloc aims to develop clean hydrogen using mainly wind and solar energy from 2030 onwards.
Financing for projects is also changing as lenders factor in the risk of unsustainable operations. The EU’s Sustainable Finance Strategy and the European Investment Bank’s impending energy lending policy (due in 2022) set stringent emission limits for fossil-based energy generation. Meanwhile the Poseidon Principles, launched in 2020, commit signatory banks to investing in that reduce emissions in line with IMO’s 2050 emission goals.
Tomorrow's worlds
With policies being put in place to spur investments in clean energy, there is little doubt that new fuels will come to market. But projecting which fuels is a difficult task. Wärtsilä explored four scenarios representing the political, social and economic factors that will have a huge effect on climate and clean energy policy and investment.
Evergreen: Sustainable fuels are in plentiful supply. A cohesive, cooperative approach to climate change between governments and industry actors trumps a fragmented approach. An idealistic perspective more than an economic one drives the development of solutions.
Silver Cloud: Sustainable fuels are supplemented by fossil fuels. Cohesive efforts are made to meet climate targets, but a financial focus means many actors are still incentivised to produce and use fossil fuels.
Red Flames: A largely fossil fuel mix is topped up with sustainable fuels. The financial justification for cleaner fuels continues to evolve, but cooperation between actors remains fragmented.
Black Fog: Idealistic resistance to the climate change means that fossil fuels remain dominant.
The Paris Agreement target is met in the Evergreen and Silver Cloud scenarios, but not in the Black Fog and Red Flames scenarios.
Each scenario was overlaid with key factors affecting fuel choices, including climate change impact, production costs, technological maturity; ease of use and public perception. The results are shown in the charts below.
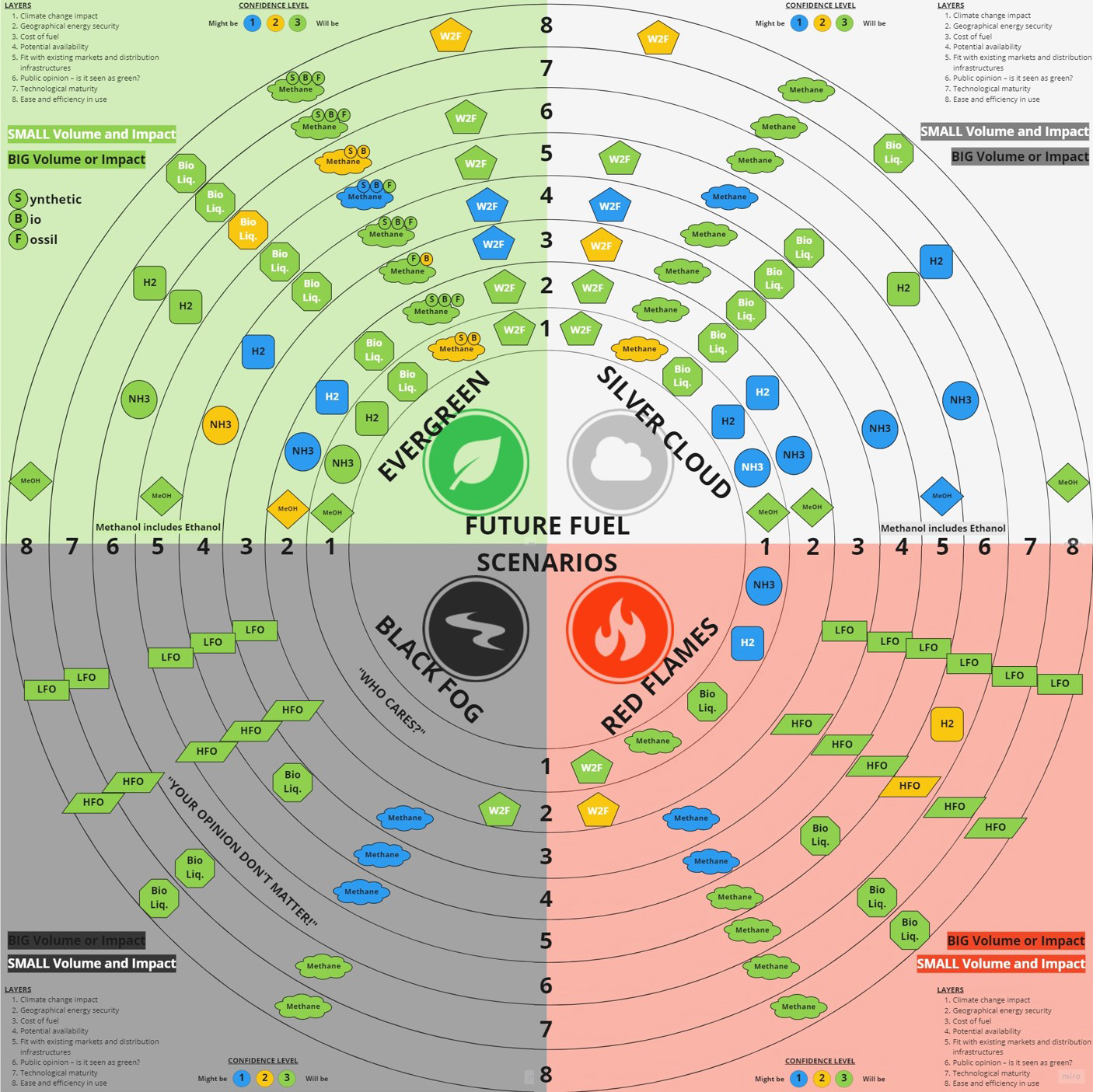

Wärtsilä examined potential fuel availability across four future scenarios
In the Evergreen and Silver Cloud scenarios, methane (produced synthetically from renewable energy or derived from biomass) is widely present, indicating a strong likelihood that it will play an important role in a successful energy transition. The other clear candidates in these scenarios include liquid biofuels and waste-to-fuel products along with renewable hydrogen and ammonia.
Sustainable fuels are less widely used in the Black Fog and Red Flames scenarios. Methane is again present, although mainly fossil generated - indicating that technology for this fuel will be an important future demand regardless of the scenario. Alongside methane, conventional fossil liquids remain the most visible fuel candidates.
Powering a cleaner planet
The critical final stage of Wärtsilä’s fuel strategy development asked which fuels are most feasible for each sector. Two example profiles in shipping highlight the difference each fuel would make (compared to marine gas oil) to annual operational costs and the annualised capital cost for technology associated with the fuel. The first is a bulk carrier with 9 MW of installed power, the second a 45 MW cruise ship. The results are shown in the two graphs below.
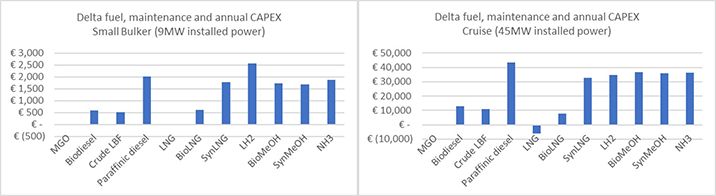
Two sample vessel profiles highlight the projected costs of using future fuels
There are some clear conclusions. Paraffinic diesel (such as hydrotreated vegetable oil) is likely too expensive for a role in shipping and best used in other industries. Bio-methanol will also probably be priced out of the sector. Other biofuels are likely to be twice the cost of fossil fuels, but would still be competitive against synthetic fuels, which incur around three times the costs associated with fossil liquids. Finally, hydrogen provides a reminder that storage and handling are as important as fuel price. The high price for storage of hydrogen negates a relatively lower fuel cost, even in the highest consumption case.
Wärtsilä used the levelized cost of electricity (LCOE) to assess the feasibility of fuels for the energy generation sector. Medium-speed engines, gas turbines and fuel cells were compared over a 20-year lifetime, with operating profiles ranging from 500 to 6,000 hours a year. The focus was on new fuels (hydrogen, ammonia, methanol) either individually or blended with the fossil natural gas or light fuel oil. The calculations exclude fuel production or storage and assume that technologies for new fuels will become less expensive as they mature.
The results for hydrogen fuel show a relatively similar LCOE across various power technologies, at between two and three times the cost of fossil fuels. This indicates that significant incentives will be needed to level the playing field with conventional fuels – a tax of around US$400 per tonne of emitted carbon dioxide equivalent would achieve cost parity with current engine technology using fuel oil.
The results are not final, says Wideskog. “There are a lot of unknowns that require further collaboration with customers. A lot of factors affecting availability and pricing could also change by the time the fuels come to market which would affect costs.”
A fuels roadmap
The preliminary findings indicate a continuing need to improve combustion technologies for fuels that are already widely used. Shipowners or plant operators using natural gas can use their existing dual-fuel or spark-ignited engines to transition to synthetic or bio methane. Diesel engines can easily switch to biodiesels and other liquid fuels. By refining these technologies further Wärtsilä can ensure that its customers retain flexibility in their fuel choices as cost and availability become clearer.
New fuels including renewably produced hydrogen, ammonia and methanol will play an important role in marine and energy markets. As these fuels are scaled up, they can be used in significant proportions as drop-in fuels to reduce the emissions footprint of existing fuels. Wärtsilä will continue to develop the enhancements to existing technologies needed for this step, as well as the more advanced engines that will be required for the full, optimised use of these fuels.
One thing is clear from Wärtsilä’s investigations. While the energy transition is in process, assets are unlikely to be operated on the same fuel for their entire lifecycle. Power plant built today may operate on natural gas, then natural gas with hydrogen, before finally running on renewable hydrogen alone. Some ships are already being built with the expectation that they will eventually switch from using LNG to ammonia. But these fuels are just two of many potential candidates. Fuel flexibility and the ability to convert for fuels will be crucial to secure uninterrupted operations independent of fuel availability while ensuring environmental performance.
This flexibility does not just apply to new vessels and power plant. Existing assets will also need upgrading in many instances. Taking an example from shipping, the IMO’s incoming Energy Efficiency Design Index for Existing Ships (EEXI) is giving shipowners another reason to consider LNG. Low-carbon fuels are one way of meeting the stringent design efficiency requirements, which are expected to come into effect from 2023. Further pressure will fall on shipowners to reduce emissions from their existing vessels as 2030 approaches. By then, IMO is aiming for a 40% reduction (compared to 2008 levels) in carbon intensity across the global fleet. Much of that reduction will need to come from vessels already in service.
As the need for fuel conversions increases, Wärtsilä’s customers can draw on long experience of successful retrofits for alternative fuels. The engine and fuel storage upgrades required for using LNG – and later synthetic or bio-methane – are already well-established technologies and have been executed across several vessel applications. Wärtsilä also has more than half a decade of experience gathered from converting diesel engines for methanol. For other fuels, including ammonia and hydrogen, solutions are under research and development.
There is little certainty around the future of fuels in either shipping or energy. But the work of the fuels steering group offers some foundations on which to build a strategy. Methane will play a role whether fossil, synthetic or from biomass. So too will hydrogen, either as a fuel or a feedstock. Flexibility will be crucial. And for many potential pathways the best solution is to develop engine and fuel handling solutions that already exist rather than create entirely new concepts. While priorities may evolve, the heart of the Wärtsilä’s fuel strategy will remain constant; to support customers by offering the highest efficiency, reliability and flexibility, whatever their fuel choices.
Did you like this? Subscribe to Insights updates!
Once every six weeks, you will get the top picks – the latest and the greatest pieces – from this Insights channel by email.



